Reducing Drug Trial Costs with Imaging Technology
Ken Tichauer
There are a number of ideas but the majority point to the assertion that the animal models used to test out drugs in preclinical trials are a poor representation of the highly variable human population that the drug is meant for.
In cancer research, this is represented by the use of human cancer cell lines that form genetically homogeneous tumors in immunosuppressed mouse models being compared to genetically heterogeneous tumors typically observed in humans (3). So, as new “genetically heterogeneous” tumor models are developed, molecular imaging approaches capable of non-invasively quantifying the heterogeneity, and the interplay of this heterogeneity with the efficacy of drug action, will be critical to the optimal evaluation of new drugs, providing:
1) an early indicator of the potential success of a drug before it goes into expensive clinical trials
2) identification of specific subpopulations of patients where the drug may or may not work (based on the molecular pathology of the cancer), to inform discretionary use of the drug
3) a means of understanding molecular reasons for a drug’s success or failure
The last decade has seen incredible advances in fluorescence and photoacoustic imaging instrumentation that have led to the development of relatively inexpensive, reproducible, and accurate tools to map the distribution of molecular targeted imaging agents three-dimensional in small animal models of disease.
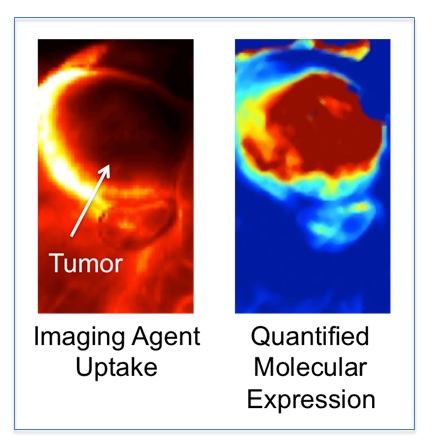
HOWEVER, mapping molecular pathology of a cancer requires more that just the ability to map the distribution of an injected molecular targeted imaging agent. Imaging agents can accumulate in or can be restricted from tumors for many physiological reasons that have nothing to do with how much of the targeted biomolecule of interest is available; e.g., blood flow, vascular permeability, interstitial pressure, enhance retention (4, 5).
The majority of the work ongoing in my lab focuses on correcting this problem of associating imaging agent uptake with targeted biomolecule expression in tumors (6, 7), with the ultimate goal of providing a more accurate tool for understanding the impact molecular pathology has on the success or failure of novel cancer therapeutics.
1. DiMasi JA, Hansen RW, & Grabowski HG (2003) The price of innovation: new estimates of drug development costs. Journal of health economics 22(2):151-185.
2. Herper M (2012) The Truly Staggering Cost of Inventing New Drugs. in PHARMA & HEALTHCARE (Forbes, http://www.forbes.com/sites/matthewherper/2012/02/10/the-truly-staggering-cost-of-inventing-new-drugs/).
3. Longo DL (2012) Tumor heterogeneity and personalized medicine. The New England journal of medicine 366(10):956-957.
4. Baeten J, Haller J, Shih H, & Ntziachristos V (2009) In vivo investigation of breast cancer progression by use of an internal control. Neoplasia 11(3):220-227.
5. Liu JT, et al. (2009) Quantifying cell-surface biomarker expression in thick tissues with ratiometric three-dimensional microscopy. Biophysical journal 96(6):2405-2414.
6. Davis SC, et al. (2013) Dynamic dual-tracer MRI-guided fluorescence tomography to quantify receptor density in vivo. Proceedings of the National Academy of Sciences of the United States of America 110(22):9025-9030.
7. Tichauer KM, et al. (2012) In vivo quantification of tumor receptor binding potential with dual-reporter molecular imaging. Molecular imaging and biology : MIB : the official publication of the Academy of Molecular Imaging 14(5):584-592.
